| repeats | diagnosis |
| 0–9 | not human |
| 10–35 | normal |
| 36–39 | high risk |
| 40–180 | Huntington's disease |
| 181–∞ | not human |
Write a program Huntingtons.java to read in a human DNA sequence from a file and identify whether the patient is at risk for HD by determining the maximum number of CAG repeats. Output the maximum number of repeats, and an appropriate medical diagnosis based on the table above. The data file format will consist of a sequence of the letters A, C, G, and T, along with arbitrary amounts of whitespace separating the letters. For full credit, your program should use regular expressions and should identify CAG repeats, even if there is intervening whitespace.
For example, the DNA sequences in test4.txt, test64.txt, chromosome4-hd.txt, and chromosome4-healthy.txt have CAG repeats of sizes 4, 64, 79, and 19, respectively.
% more test4.txt % more test64.txt
TTTTTTTTTT TTTTTTTTTT TTTTTTTTCA TTTTTTTTTT TTTTTTTTTT TTTTTTTTTT
GCAGCAGCAG TTTCAGCAGT TTTTTTTTTT TTTTTTTTTT TTTTTTTTTT TTCAGCAGCA
TTTTTTTTTT TTTTTTTTTT TTTTTTTTTT GCAGCAGCAG CAGCAGCAGC AGCAGCAGCA
TTTCAGTTTT TTTTTTTTTT T GCAGCAGCAG CAGCAGCAGC AGCAGCAGCA
GCAGCAGCAG CAGCAGCAGC AGCAGCAGCA
GCAGCAGCAG CAGCAGCAGC AGCAGCAGCA
GCAGCAGCAG CAGCAGCAGC AGCAGCAGCA
GCAGCAGCAG CAGCAGCAGC AGCAGCAGCA
GCAGTTTTTT TTTTTTTTTT TTTTTTTTTT
TTTTTTTTTT TTTTTTTTTT TTTTTTTTTT
% java Huntingtons test4.txt %java Huntingtons test64.txt
max number of CAG repeats = 4 max number of CAG repeats = 64
not human Huntington's disease
% java Huntingtons chromosome4-hd.txt % java Huntingtons chromosome4-healthy.txt
max number of CAG repeats = 79 max number of CAG repeats = 19
Huntington's disease normal
Use this Turing machine simulator to develop your machine. Your job is to create and submit a text file comparator.tur containing the description of your Turing machine like the one below, but instead of adding the two binary numbers, it should compare them.
title Binary Adder description K. Wayne, wayne, P01 Add two binary numbers, and leave the result on the tape. vertices 2 R 0.25 0.75 0 L 0.25 0.25 1 L 0.75 0.25 3 L 0.75 0.75 4 R 0.40 0.50 5 H 0.60 0.50 edges 0 0 0 1 135 0 1 1 0 0 4 + # 1 3 + + 2 0 # # 3 2 # 1 0.2 3 2 0 1 -0.2 3 3 1 0 -30 4 4 1 # -135 4 5 # # tapes [1] 0 1 0 + 1 1 1 1 [1] 0 0 + 1 1 1 [1] 1 0 0 + 1 1 [1] 0 1 0 + 1 0 0 1 0 1 |
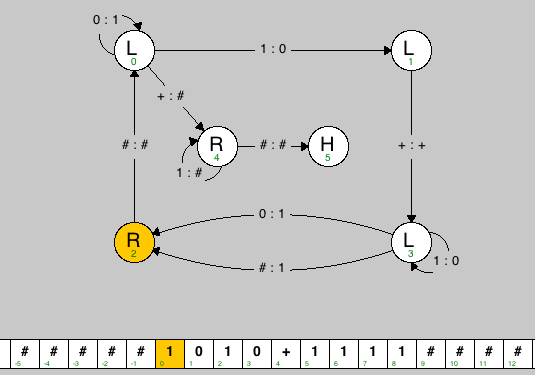 |
The data file is divided into 5 sections (title, description, vertices, edges, tapes), and each section begins with the corresponding name. Each vertex (state) is specified with 4 required values: its name (a string), its type (L, R, Y, N, H), and its x- and y-coordinates (between 0 and 1). The first state listed is the start state. Each edge (transition) is specified with 4 required values: the starting state, the ending state, the symbol to read, and the symbol to write. An optional 5th parameter specifies the curvature with which to draw the edge (straight line by default). Each sample input tape is specified by listing its symbols, separated by spaces. The tape head starts at the symbol delimited by square braces - for this assignment, it should always be the leftmost non-blank character.
Getting started. The subdirectory theory from the COS126 ftp site contains Grep.java, Harvester.java, In.java and StdOut.java as well as several small test files and two larger files with actual chromosome data for your program for Part 1. There is also the example Turing Machine file, binaryadder.tur. As always, there is a readme.txt for you to complete.
Submission. Submit Huntingtons.java, comparator.tur, and readme.txt.
Extra credit. If you based your comparator on binaryadder.tur, then it is very similar to a subtractor. Unfortunately, it would take more than 2n steps to subtract two n-bit numbers. Your challenge is to design a Turing machine that subtracts two positive n-bit numbers in time proportional to n2. Assume the left number on the tape is always larger than or equal to the right number, so the difference will always be greater than or equal to zero. However, do not assume that the two numbers always have the same number of bits. Call your file efficientSubtractor.tur and submit it as an Additional File.
This assignment was created by Kevin Wayne. Copyright © 2004.